CATEGORY
Molecular Properties
SOURCE
RDKit
DESCRIPTION
Generate 3D conformers of molecules using RDKit
INPUTS
A Dataset of molecules
OUTPUTS
A Dataset of 3D molecules
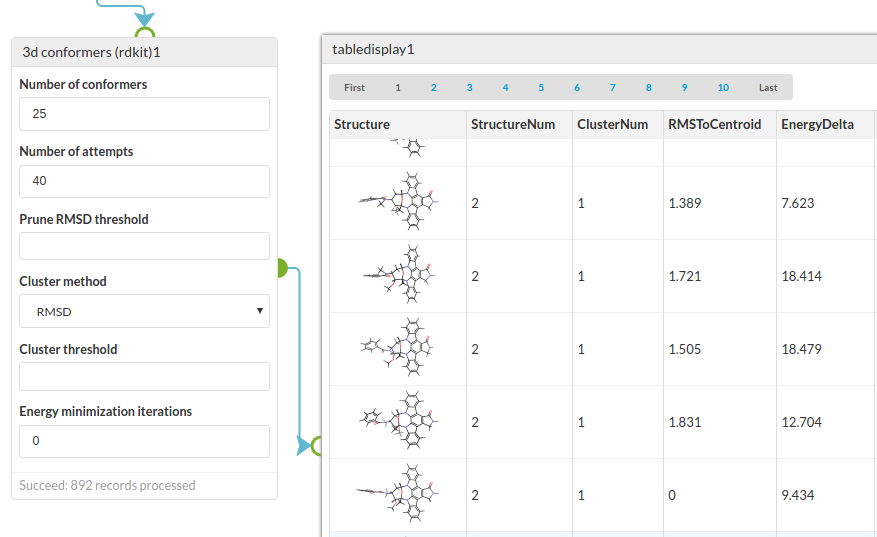
OPTIONS
| Number of conformers | The desired number of conformers to generate (param num below). |
| Number of attempts | The maximum number of attempts used to generate the conformers for a molecule. Must be not be less than the number of conformers. If not specified then the number of attempts is the number of requested conformers (param attempts below). |
| Prune RMSD threshold | Exclude a conformer that is within this RMSD of an existing conformer (param prune below) |
| Cluster method | How to cluster the molecules. Using Root Means Squared Deviation (RMSD) or Torsion Fingerprint Deviation (TFD) (param method below). |
| Cluster threshold | Threshold for clustering. The value used depends on the cluster method. If not specified the defaults are 2.0 for RMSD and 0.3 for TFD (param thresh below). |
| Energy minimisation iterations | The number of iterations of energy minimisation using the MMFF. Default is zero. See note below (param minimise below). |
These parameters are described in more detail below.
ADDITIONAL INFO
This cell implements a fairly sophisticated conformer generation algorithm. using RDKit. More about the 3D capabilities of RDKit can be found in the RDKit cookbook.
The approach used is as follows:
For each molecule in the input:
- Attempt to generate num conformers. Very similar conformers (as determined by an RMSD of less than prune to an existing conformer) are removed. Up to attempts conformers are tried to generate the specified num of conformers. If similar conformers are found then it might not be possible to reach num. The value of attempts must be equal to, or greater than num.
- For each conformer optionally perform minimise energy minimisation iterations on each structure using the Merck Molecular Force Field (MMFF) and then calculate the energy. See below about the need for energy minimisation.
- The generated conformers are clustered using Butina clustering using the specified method. RMSD uses the differences in the atom coordinates, TFD uses the differences in torsion angles. For more information of TFD look at Schulz-Gasch et al. (J. Chem. Inf. Model., 52, 1499, 2012)
- For each cluster calculate the RMSD and energy difference of each conformer to the cluster centroid.
- Write out the resulting conformers and properties.
RDKit conformer generation
RDKit uses a distance geometry embedding approach. Recently this has been enhanced with a technique named ETKDG that uses a knowledge based approach to generate reasonable structures (e.g. aromatic rings are flat). This means that energy minimisation is not normally needed to generate “good” structures, as would normally be the case with distance geometry embedding approaches. We provide the option of doing energy minimisations, but by default this is turned off (minimise = 0) which should result in better sampling of conformational space as minimisation does not drag conformations down to the same low energy conformations. If however, you specifically want lowest energy conformations as opposed for good sampling then you can specify a number of minimise iterations.
The ETKDG approach is described in Riniker & Landrum (J. Chem. Inf. Model., 2015, 55 (12), pp 2562–2574).