Running jobs - a How To guide
Use the Apps/Jobs tab to launch Jobs to do your work. Applications are also listed here - see the Applications How To guide for details of launching these.
The jobs that are available (defined by the system administrator) are displayed as "cards". You can type into the "Search" box to filter these using terms that are defined for each job or app. For instance jobs implemented using RDKit should be labelled as "rdkit" so typing "rdkit" into the search box will show only cards that have this label. The labels for each job are displayed towards the bottom of the card in red.
Each job belongs to a category (the type of job it is e.g. molecular properties) and a collection (where it comes from e.g. rdkit). A link the the job's documentation is also present, providing more information about the job and how to run it.
To run a job or application click on the Run button. A dialog will appear allowing you to define the job inputs and options.
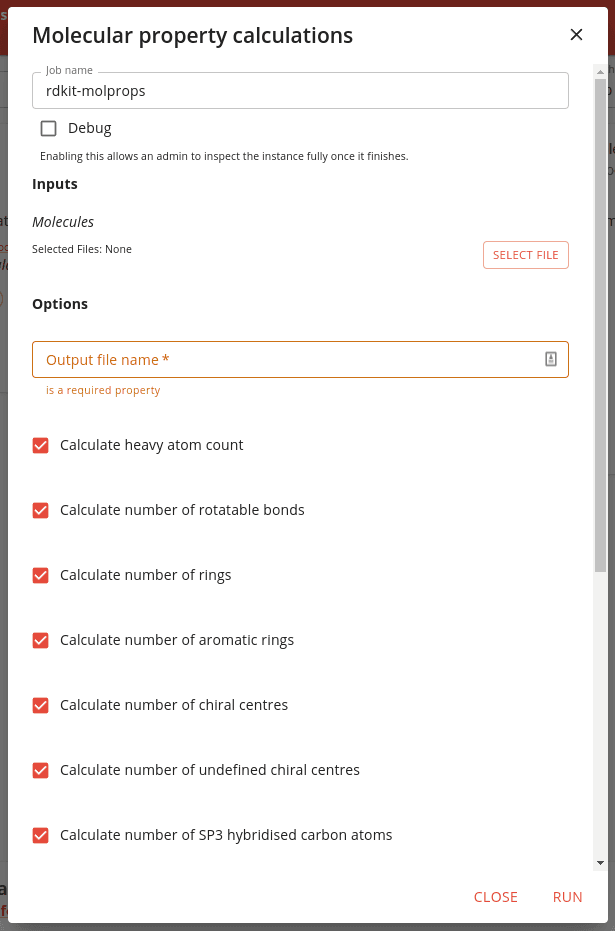
Inputs are project files that form the input to the job.
Options are user specified parameters for execution of the
job e.g. which properties to calculate in the example above. An important option is often the name of the output file.
We recommend putting the outputs of a job in a new subdirectory so that the output of one job does not overwrite that of
another.
Most jobs are implemented so that you can specify a full path to the file you want, including subdirectories. e.g. you
can specify some/where/results.sdf as the output file name and the directories some/where/ will be created for you
(from the project root).
Specify these inputs and options and click the Run button at the bottom. The execution will be added to the bottom of the card. Click on it to see the execution details in the Results tab. See the Inspecting job and application execution How To guide for details.
